Transcript Detail IRES Map
Queried Transcript
| Transcript Info | Conservation | Conditional Translation Efficiency | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Accession | Genome Location |
Trascribed Gene |
5'UTR | CDS | Stop Codon | 3'UTR | Average Phastcon Score |
ER stressed | Normal |
ER/Normal Ratio |
| NM_001256621 | chr10: + (location) |
R3HCC1L |
245 bps (seq) |
552 bps (seq) |
798 ..801 (TGA) |
739 bps (seq) |
0.707 | 0.000 | 0.000 |
NaN
(q = NaN) |
Identified Putative IRES Elements
* indicates statistical significance:
(Thresholds
selected on a ground truth dataset)
1. Conservation Score q-value < 1.000E-6
2. RPI Score q-value < 1.000E-6.
3. Ratio (difference) q-value < 1.000E-6.
| IRES Info | Conservation |
Conditional
Translation Efficiency |
Translation Initiation-related Info Summary for the IRES | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Putative IRES | Identifying Tools |
Average Phastcon Score (ref) |
ER Stressed | Normal | ER/Normal Ratio (ref) |
# of eTIS
(ref) |
# of nTIS
(ref) |
# of uORF
(ref) |
# of ITAF Binding Events (ref) |
RPI Significance |
IRES activity (ref) |
# of Literature Evidence |
|
IRES 1 149..245 (seq) (location) |
UTRscan |
0.186
(q = 1.000) |
0.000 | 0.000 |
NaN
(q = NaN) |
0 | 0 | 3 | 3 | q = 2.833E-13* | -- | 0 |
IRES-Translation Initiation Interaction Map
(TIS refers to the main ORF initiating codon)
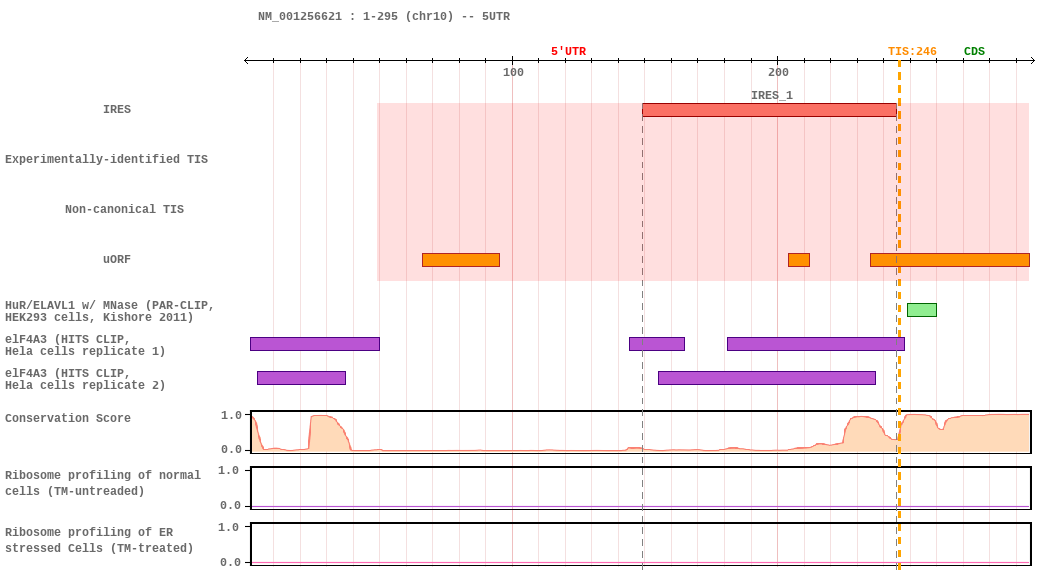
Detailed Translation Initiation-Related Data
Phastcon Conservation Scores (Among 19 Mammalian Species)
| Interconnected IRES |
Transcript Location |
Chromosomal Location |
Conservation Score |
|---|---|---|---|
| IRES 1 (149..245) | 149 | chr10 (+): 98163298 | 0.059 |
| IRES 1 (149..245) | 150 | chr10 (+): 98163299 | 0.032 |
| IRES 1 (149..245) | 151 | chr10 (+): 98163300 | 0.029 |
| IRES 1 (149..245) | 152 | chr10 (+): 98163301 | 0.018 |
| IRES 1 (149..245) | 153 | chr10 (+): 98163302 | 0.010 |
| IRES 1 (149..245) | 154 | chr10 (+): 98163303 | 0.006 |
| IRES 1 (149..245) | 155 | chr10 (+): 98163304 | 0.003 |
| IRES 1 (149..245) | 156 | chr10 (+): 98163305 | 0.002 |
| IRES 1 (149..245) | 157 | chr10 (+): 98163306 | 0.000 |
| IRES 1 (149..245) | 158 | chr10 (+): 98163307 | 0.006 |
| IRES 1 (149..245) | 159 | chr10 (+): 98163308 | 0.009 |
| IRES 1 (149..245) | 160 | chr10 (+): 98163309 | 0.013 |
| IRES 1 (149..245) | 161 | chr10 (+): 98163310 | 0.014 |
| IRES 1 (149..245) | 162 | chr10 (+): 98163311 | 0.011 |
| IRES 1 (149..245) | 163 | chr10 (+): 98163312 | 0.013 |
| IRES 1 (149..245) | 164 | chr10 (+): 98163313 | 0.012 |
| IRES 1 (149..245) | 165 | chr10 (+): 98163314 | 0.012 |
| IRES 1 (149..245) | 166 | chr10 (+): 98163315 | 0.009 |
| IRES 1 (149..245) | 167 | chr10 (+): 98163316 | 0.010 |
| IRES 1 (149..245) | 168 | chr10 (+): 98163317 | 0.013 |
| IRES 1 (149..245) | 169 | chr10 (+): 98163318 | 0.016 |
| IRES 1 (149..245) | 170 | chr10 (+): 98163319 | 0.017 |
| IRES 1 (149..245) | 171 | chr10 (+): 98163320 | 0.014 |
| IRES 1 (149..245) | 172 | chr10 (+): 98163321 | 0.006 |
| IRES 1 (149..245) | 173 | chr10 (+): 98163322 | 0.001 |
| IRES 1 (149..245) | 174 | chr10 (+): 98163323 | 0.001 |
| IRES 1 (149..245) | 175 | chr10 (+): 98163324 | 0.001 |
| IRES 1 (149..245) | 176 | chr10 (+): 98163325 | 0.001 |
| IRES 1 (149..245) | 177 | chr10 (+): 98163326 | 0.002 |
| IRES 1 (149..245) | 178 | chr10 (+): 98163327 | 0.005 |
| IRES 1 (149..245) | 179 | chr10 (+): 98163328 | 0.026 |
| IRES 1 (149..245) | 180 | chr10 (+): 98163329 | 0.034 |
| IRES 1 (149..245) | 181 | chr10 (+): 98163330 | 0.045 |
| IRES 1 (149..245) | 182 | chr10 (+): 98163331 | 0.062 |
| IRES 1 (149..245) | 183 | chr10 (+): 98163332 | 0.070 |
| IRES 1 (149..245) | 184 | chr10 (+): 98163333 | 0.071 |
| IRES 1 (149..245) | 185 | chr10 (+): 98163334 | 0.065 |
| IRES 1 (149..245) | 186 | chr10 (+): 98163335 | 0.047 |
| IRES 1 (149..245) | 187 | chr10 (+): 98163336 | 0.047 |
| IRES 1 (149..245) | 188 | chr10 (+): 98163337 | 0.041 |
| IRES 1 (149..245) | 189 | chr10 (+): 98163338 | 0.023 |
| IRES 1 (149..245) | 190 | chr10 (+): 98163339 | 0.011 |
| IRES 1 (149..245) | 191 | chr10 (+): 98163340 | 0.010 |
| IRES 1 (149..245) | 192 | chr10 (+): 98163341 | 0.004 |
| IRES 1 (149..245) | 193 | chr10 (+): 98163342 | 0.004 |
| IRES 1 (149..245) | 194 | chr10 (+): 98163343 | 0.002 |
| IRES 1 (149..245) | 195 | chr10 (+): 98163344 | 0.002 |
| IRES 1 (149..245) | 196 | chr10 (+): 98163345 | 0.001 |
| IRES 1 (149..245) | 197 | chr10 (+): 98163346 | 0.002 |
| IRES 1 (149..245) | 198 | chr10 (+): 98163347 | 0.002 |
| IRES 1 (149..245) | 199 | chr10 (+): 98163348 | 0.006 |
| IRES 1 (149..245) | 200 | chr10 (+): 98163349 | 0.007 |
| IRES 1 (149..245) | 201 | chr10 (+): 98163350 | 0.005 |
| IRES 1 (149..245) | 202 | chr10 (+): 98163351 | 0.007 |
| IRES 1 (149..245) | 203 | chr10 (+): 98163352 | 0.007 |
| IRES 1 (149..245) | 204 | chr10 (+): 98163353 | 0.007 |
| IRES 1 (149..245) | 205 | chr10 (+): 98163354 | 0.034 |
| IRES 1 (149..245) | 206 | chr10 (+): 98163355 | 0.042 |
| IRES 1 (149..245) | 207 | chr10 (+): 98163356 | 0.054 |
| IRES 1 (149..245) | 208 | chr10 (+): 98163357 | 0.066 |
| IRES 1 (149..245) | 209 | chr10 (+): 98163358 | 0.068 |
| IRES 1 (149..245) | 210 | chr10 (+): 98163359 | 0.071 |
| IRES 1 (149..245) | 211 | chr10 (+): 98163360 | 0.075 |
| IRES 1 (149..245) | 212 | chr10 (+): 98163361 | 0.080 |
| IRES 1 (149..245) | 213 | chr10 (+): 98163362 | 0.087 |
| IRES 1 (149..245) | 214 | chr10 (+): 98163363 | 0.127 |
| IRES 1 (149..245) | 215 | chr10 (+): 98163364 | 0.177 |
| IRES 1 (149..245) | 216 | chr10 (+): 98163365 | 0.192 |
| IRES 1 (149..245) | 217 | chr10 (+): 98163366 | 0.191 |
| IRES 1 (149..245) | 218 | chr10 (+): 98163367 | 0.175 |
| IRES 1 (149..245) | 219 | chr10 (+): 98163368 | 0.162 |
| IRES 1 (149..245) | 220 | chr10 (+): 98163369 | 0.152 |
| IRES 1 (149..245) | 221 | chr10 (+): 98163370 | 0.159 |
| IRES 1 (149..245) | 222 | chr10 (+): 98163371 | 0.169 |
| IRES 1 (149..245) | 223 | chr10 (+): 98163372 | 0.184 |
| IRES 1 (149..245) | 224 | chr10 (+): 98163373 | 0.202 |
| IRES 1 (149..245) | 225 | chr10 (+): 98163374 | 0.207 |
| IRES 1 (149..245) | 226 | chr10 (+): 98163375 | 0.600 |
| IRES 1 (149..245) | 227 | chr10 (+): 98163376 | 0.729 |
| IRES 1 (149..245) | 228 | chr10 (+): 98163377 | 0.878 |
| IRES 1 (149..245) | 229 | chr10 (+): 98163378 | 0.926 |
| IRES 1 (149..245) | 230 | chr10 (+): 98163379 | 0.942 |
| IRES 1 (149..245) | 231 | chr10 (+): 98163380 | 0.945 |
| IRES 1 (149..245) | 232 | chr10 (+): 98163381 | 0.950 |
| IRES 1 (149..245) | 233 | chr10 (+): 98163382 | 0.947 |
| IRES 1 (149..245) | 234 | chr10 (+): 98163383 | 0.935 |
| IRES 1 (149..245) | 235 | chr10 (+): 98163384 | 0.925 |
| IRES 1 (149..245) | 236 | chr10 (+): 98163385 | 0.892 |
| IRES 1 (149..245) | 237 | chr10 (+): 98163386 | 0.868 |
| IRES 1 (149..245) | 238 | chr10 (+): 98163387 | 0.812 |
| IRES 1 (149..245) | 239 | chr10 (+): 98163388 | 0.680 |
| IRES 1 (149..245) | 240 | chr10 (+): 98163389 | 0.607 |
| IRES 1 (149..245) | 241 | chr10 (+): 98163390 | 0.429 |
| IRES 1 (149..245) | 242 | chr10 (+): 98163391 | 0.403 |
| IRES 1 (149..245) | 243 | chr10 (+): 98163392 | 0.340 |
| IRES 1 (149..245) | 244 | chr10 (+): 98163393 | 0.298 |
| IRES 1 (149..245) | 245 | chr10 (+): 98163394 | 0.294 |
Potential IRES Structural Form
IRES 1
(149.. 245)
Best Predicted IRES Structural Form:
(((((((((.(.((((((((.......))))..)))))...)))))).....))).....((((.((((.(((........))))))).))))....
Predicted by: RNAprob
Structural RPI Interpretability Significance: q = 2.833E-13*
(Visualization)
Conditional Translation Activity
ER-Stressed Ribisome Profiling (TM-treated)
| Interconnected IRES |
Transcript Location |
Chromosomal Location |
Stressed Translation Activity |
|---|---|---|---|
| IRES 1 (149..245) | 149 | chr10 (+): 98163298 | 0 |
| IRES 1 (149..245) | 150 | chr10 (+): 98163299 | 0 |
| IRES 1 (149..245) | 151 | chr10 (+): 98163300 | 0 |
| IRES 1 (149..245) | 152 | chr10 (+): 98163301 | 0 |
| IRES 1 (149..245) | 153 | chr10 (+): 98163302 | 0 |
| IRES 1 (149..245) | 154 | chr10 (+): 98163303 | 0 |
| IRES 1 (149..245) | 155 | chr10 (+): 98163304 | 0 |
| IRES 1 (149..245) | 156 | chr10 (+): 98163305 | 0 |
| IRES 1 (149..245) | 157 | chr10 (+): 98163306 | 0 |
| IRES 1 (149..245) | 158 | chr10 (+): 98163307 | 0 |
| IRES 1 (149..245) | 159 | chr10 (+): 98163308 | 0 |
| IRES 1 (149..245) | 160 | chr10 (+): 98163309 | 0 |
| IRES 1 (149..245) | 161 | chr10 (+): 98163310 | 0 |
| IRES 1 (149..245) | 162 | chr10 (+): 98163311 | 0 |
| IRES 1 (149..245) | 163 | chr10 (+): 98163312 | 0 |
| IRES 1 (149..245) | 164 | chr10 (+): 98163313 | 0 |
| IRES 1 (149..245) | 165 | chr10 (+): 98163314 | 0 |
| IRES 1 (149..245) | 166 | chr10 (+): 98163315 | 0 |
| IRES 1 (149..245) | 167 | chr10 (+): 98163316 | 0 |
| IRES 1 (149..245) | 168 | chr10 (+): 98163317 | 0 |
| IRES 1 (149..245) | 169 | chr10 (+): 98163318 | 0 |
| IRES 1 (149..245) | 170 | chr10 (+): 98163319 | 0 |
| IRES 1 (149..245) | 171 | chr10 (+): 98163320 | 0 |
| IRES 1 (149..245) | 172 | chr10 (+): 98163321 | 0 |
| IRES 1 (149..245) | 173 | chr10 (+): 98163322 | 0 |
| IRES 1 (149..245) | 174 | chr10 (+): 98163323 | 0 |
| IRES 1 (149..245) | 175 | chr10 (+): 98163324 | 0 |
| IRES 1 (149..245) | 176 | chr10 (+): 98163325 | 0 |
| IRES 1 (149..245) | 177 | chr10 (+): 98163326 | 0 |
| IRES 1 (149..245) | 178 | chr10 (+): 98163327 | 0 |
| IRES 1 (149..245) | 179 | chr10 (+): 98163328 | 0 |
| IRES 1 (149..245) | 180 | chr10 (+): 98163329 | 0 |
| IRES 1 (149..245) | 181 | chr10 (+): 98163330 | 0 |
| IRES 1 (149..245) | 182 | chr10 (+): 98163331 | 0 |
| IRES 1 (149..245) | 183 | chr10 (+): 98163332 | 0 |
| IRES 1 (149..245) | 184 | chr10 (+): 98163333 | 0 |
| IRES 1 (149..245) | 185 | chr10 (+): 98163334 | 0 |
| IRES 1 (149..245) | 186 | chr10 (+): 98163335 | 0 |
| IRES 1 (149..245) | 187 | chr10 (+): 98163336 | 0 |
| IRES 1 (149..245) | 188 | chr10 (+): 98163337 | 0 |
| IRES 1 (149..245) | 189 | chr10 (+): 98163338 | 0 |
| IRES 1 (149..245) | 190 | chr10 (+): 98163339 | 0 |
| IRES 1 (149..245) | 191 | chr10 (+): 98163340 | 0 |
| IRES 1 (149..245) | 192 | chr10 (+): 98163341 | 0 |
| IRES 1 (149..245) | 193 | chr10 (+): 98163342 | 0 |
| IRES 1 (149..245) | 194 | chr10 (+): 98163343 | 0 |
| IRES 1 (149..245) | 195 | chr10 (+): 98163344 | 0 |
| IRES 1 (149..245) | 196 | chr10 (+): 98163345 | 0 |
| IRES 1 (149..245) | 197 | chr10 (+): 98163346 | 0 |
| IRES 1 (149..245) | 198 | chr10 (+): 98163347 | 0 |
| IRES 1 (149..245) | 199 | chr10 (+): 98163348 | 0 |
| IRES 1 (149..245) | 200 | chr10 (+): 98163349 | 0 |
| IRES 1 (149..245) | 201 | chr10 (+): 98163350 | 0 |
| IRES 1 (149..245) | 202 | chr10 (+): 98163351 | 0 |
| IRES 1 (149..245) | 203 | chr10 (+): 98163352 | 0 |
| IRES 1 (149..245) | 204 | chr10 (+): 98163353 | 0 |
| IRES 1 (149..245) | 205 | chr10 (+): 98163354 | 0 |
| IRES 1 (149..245) | 206 | chr10 (+): 98163355 | 0 |
| IRES 1 (149..245) | 207 | chr10 (+): 98163356 | 0 |
| IRES 1 (149..245) | 208 | chr10 (+): 98163357 | 0 |
| IRES 1 (149..245) | 209 | chr10 (+): 98163358 | 0 |
| IRES 1 (149..245) | 210 | chr10 (+): 98163359 | 0 |
| IRES 1 (149..245) | 211 | chr10 (+): 98163360 | 0 |
| IRES 1 (149..245) | 212 | chr10 (+): 98163361 | 0 |
| IRES 1 (149..245) | 213 | chr10 (+): 98163362 | 0 |
| IRES 1 (149..245) | 214 | chr10 (+): 98163363 | 0 |
| IRES 1 (149..245) | 215 | chr10 (+): 98163364 | 0 |
| IRES 1 (149..245) | 216 | chr10 (+): 98163365 | 0 |
| IRES 1 (149..245) | 217 | chr10 (+): 98163366 | 0 |
| IRES 1 (149..245) | 218 | chr10 (+): 98163367 | 0 |
| IRES 1 (149..245) | 219 | chr10 (+): 98163368 | 0 |
| IRES 1 (149..245) | 220 | chr10 (+): 98163369 | 0 |
| IRES 1 (149..245) | 221 | chr10 (+): 98163370 | 0 |
| IRES 1 (149..245) | 222 | chr10 (+): 98163371 | 0 |
| IRES 1 (149..245) | 223 | chr10 (+): 98163372 | 0 |
| IRES 1 (149..245) | 224 | chr10 (+): 98163373 | 0 |
| IRES 1 (149..245) | 225 | chr10 (+): 98163374 | 0 |
| IRES 1 (149..245) | 226 | chr10 (+): 98163375 | 0 |
| IRES 1 (149..245) | 227 | chr10 (+): 98163376 | 0 |
| IRES 1 (149..245) | 228 | chr10 (+): 98163377 | 0 |
| IRES 1 (149..245) | 229 | chr10 (+): 98163378 | 0 |
| IRES 1 (149..245) | 230 | chr10 (+): 98163379 | 0 |
| IRES 1 (149..245) | 231 | chr10 (+): 98163380 | 0 |
| IRES 1 (149..245) | 232 | chr10 (+): 98163381 | 0 |
| IRES 1 (149..245) | 233 | chr10 (+): 98163382 | 0 |
| IRES 1 (149..245) | 234 | chr10 (+): 98163383 | 0 |
| IRES 1 (149..245) | 235 | chr10 (+): 98163384 | 0 |
| IRES 1 (149..245) | 236 | chr10 (+): 98163385 | 0 |
| IRES 1 (149..245) | 237 | chr10 (+): 98163386 | 0 |
| IRES 1 (149..245) | 238 | chr10 (+): 98163387 | 0 |
| IRES 1 (149..245) | 239 | chr10 (+): 98163388 | 0 |
| IRES 1 (149..245) | 240 | chr10 (+): 98163389 | 0 |
| IRES 1 (149..245) | 241 | chr10 (+): 98163390 | 0 |
| IRES 1 (149..245) | 242 | chr10 (+): 98163391 | 0 |
| IRES 1 (149..245) | 243 | chr10 (+): 98163392 | 0 |
| IRES 1 (149..245) | 244 | chr10 (+): 98163393 | 0 |
| IRES 1 (149..245) | 245 | chr10 (+): 98163394 | 0 |
Normal Ribisome Profiling (TM-untreaed)
| Interconnected IRES |
Transcript Location |
Chromosomal Location |
Normal Translation Activity |
|---|---|---|---|
| IRES 1 (149..245) | 149 | chr10 (+): 98163298 | 0 |
| IRES 1 (149..245) | 150 | chr10 (+): 98163299 | 0 |
| IRES 1 (149..245) | 151 | chr10 (+): 98163300 | 0 |
| IRES 1 (149..245) | 152 | chr10 (+): 98163301 | 0 |
| IRES 1 (149..245) | 153 | chr10 (+): 98163302 | 0 |
| IRES 1 (149..245) | 154 | chr10 (+): 98163303 | 0 |
| IRES 1 (149..245) | 155 | chr10 (+): 98163304 | 0 |
| IRES 1 (149..245) | 156 | chr10 (+): 98163305 | 0 |
| IRES 1 (149..245) | 157 | chr10 (+): 98163306 | 0 |
| IRES 1 (149..245) | 158 | chr10 (+): 98163307 | 0 |
| IRES 1 (149..245) | 159 | chr10 (+): 98163308 | 0 |
| IRES 1 (149..245) | 160 | chr10 (+): 98163309 | 0 |
| IRES 1 (149..245) | 161 | chr10 (+): 98163310 | 0 |
| IRES 1 (149..245) | 162 | chr10 (+): 98163311 | 0 |
| IRES 1 (149..245) | 163 | chr10 (+): 98163312 | 0 |
| IRES 1 (149..245) | 164 | chr10 (+): 98163313 | 0 |
| IRES 1 (149..245) | 165 | chr10 (+): 98163314 | 0 |
| IRES 1 (149..245) | 166 | chr10 (+): 98163315 | 0 |
| IRES 1 (149..245) | 167 | chr10 (+): 98163316 | 0 |
| IRES 1 (149..245) | 168 | chr10 (+): 98163317 | 0 |
| IRES 1 (149..245) | 169 | chr10 (+): 98163318 | 0 |
| IRES 1 (149..245) | 170 | chr10 (+): 98163319 | 0 |
| IRES 1 (149..245) | 171 | chr10 (+): 98163320 | 0 |
| IRES 1 (149..245) | 172 | chr10 (+): 98163321 | 0 |
| IRES 1 (149..245) | 173 | chr10 (+): 98163322 | 0 |
| IRES 1 (149..245) | 174 | chr10 (+): 98163323 | 0 |
| IRES 1 (149..245) | 175 | chr10 (+): 98163324 | 0 |
| IRES 1 (149..245) | 176 | chr10 (+): 98163325 | 0 |
| IRES 1 (149..245) | 177 | chr10 (+): 98163326 | 0 |
| IRES 1 (149..245) | 178 | chr10 (+): 98163327 | 0 |
| IRES 1 (149..245) | 179 | chr10 (+): 98163328 | 0 |
| IRES 1 (149..245) | 180 | chr10 (+): 98163329 | 0 |
| IRES 1 (149..245) | 181 | chr10 (+): 98163330 | 0 |
| IRES 1 (149..245) | 182 | chr10 (+): 98163331 | 0 |
| IRES 1 (149..245) | 183 | chr10 (+): 98163332 | 0 |
| IRES 1 (149..245) | 184 | chr10 (+): 98163333 | 0 |
| IRES 1 (149..245) | 185 | chr10 (+): 98163334 | 0 |
| IRES 1 (149..245) | 186 | chr10 (+): 98163335 | 0 |
| IRES 1 (149..245) | 187 | chr10 (+): 98163336 | 0 |
| IRES 1 (149..245) | 188 | chr10 (+): 98163337 | 0 |
| IRES 1 (149..245) | 189 | chr10 (+): 98163338 | 0 |
| IRES 1 (149..245) | 190 | chr10 (+): 98163339 | 0 |
| IRES 1 (149..245) | 191 | chr10 (+): 98163340 | 0 |
| IRES 1 (149..245) | 192 | chr10 (+): 98163341 | 0 |
| IRES 1 (149..245) | 193 | chr10 (+): 98163342 | 0 |
| IRES 1 (149..245) | 194 | chr10 (+): 98163343 | 0 |
| IRES 1 (149..245) | 195 | chr10 (+): 98163344 | 0 |
| IRES 1 (149..245) | 196 | chr10 (+): 98163345 | 0 |
| IRES 1 (149..245) | 197 | chr10 (+): 98163346 | 0 |
| IRES 1 (149..245) | 198 | chr10 (+): 98163347 | 0 |
| IRES 1 (149..245) | 199 | chr10 (+): 98163348 | 0 |
| IRES 1 (149..245) | 200 | chr10 (+): 98163349 | 0 |
| IRES 1 (149..245) | 201 | chr10 (+): 98163350 | 0 |
| IRES 1 (149..245) | 202 | chr10 (+): 98163351 | 0 |
| IRES 1 (149..245) | 203 | chr10 (+): 98163352 | 0 |
| IRES 1 (149..245) | 204 | chr10 (+): 98163353 | 0 |
| IRES 1 (149..245) | 205 | chr10 (+): 98163354 | 0 |
| IRES 1 (149..245) | 206 | chr10 (+): 98163355 | 0 |
| IRES 1 (149..245) | 207 | chr10 (+): 98163356 | 0 |
| IRES 1 (149..245) | 208 | chr10 (+): 98163357 | 0 |
| IRES 1 (149..245) | 209 | chr10 (+): 98163358 | 0 |
| IRES 1 (149..245) | 210 | chr10 (+): 98163359 | 0 |
| IRES 1 (149..245) | 211 | chr10 (+): 98163360 | 0 |
| IRES 1 (149..245) | 212 | chr10 (+): 98163361 | 0 |
| IRES 1 (149..245) | 213 | chr10 (+): 98163362 | 0 |
| IRES 1 (149..245) | 214 | chr10 (+): 98163363 | 0 |
| IRES 1 (149..245) | 215 | chr10 (+): 98163364 | 0 |
| IRES 1 (149..245) | 216 | chr10 (+): 98163365 | 0 |
| IRES 1 (149..245) | 217 | chr10 (+): 98163366 | 0 |
| IRES 1 (149..245) | 218 | chr10 (+): 98163367 | 0 |
| IRES 1 (149..245) | 219 | chr10 (+): 98163368 | 0 |
| IRES 1 (149..245) | 220 | chr10 (+): 98163369 | 0 |
| IRES 1 (149..245) | 221 | chr10 (+): 98163370 | 0 |
| IRES 1 (149..245) | 222 | chr10 (+): 98163371 | 0 |
| IRES 1 (149..245) | 223 | chr10 (+): 98163372 | 0 |
| IRES 1 (149..245) | 224 | chr10 (+): 98163373 | 0 |
| IRES 1 (149..245) | 225 | chr10 (+): 98163374 | 0 |
| IRES 1 (149..245) | 226 | chr10 (+): 98163375 | 0 |
| IRES 1 (149..245) | 227 | chr10 (+): 98163376 | 0 |
| IRES 1 (149..245) | 228 | chr10 (+): 98163377 | 0 |
| IRES 1 (149..245) | 229 | chr10 (+): 98163378 | 0 |
| IRES 1 (149..245) | 230 | chr10 (+): 98163379 | 0 |
| IRES 1 (149..245) | 231 | chr10 (+): 98163380 | 0 |
| IRES 1 (149..245) | 232 | chr10 (+): 98163381 | 0 |
| IRES 1 (149..245) | 233 | chr10 (+): 98163382 | 0 |
| IRES 1 (149..245) | 234 | chr10 (+): 98163383 | 0 |
| IRES 1 (149..245) | 235 | chr10 (+): 98163384 | 0 |
| IRES 1 (149..245) | 236 | chr10 (+): 98163385 | 0 |
| IRES 1 (149..245) | 237 | chr10 (+): 98163386 | 0 |
| IRES 1 (149..245) | 238 | chr10 (+): 98163387 | 0 |
| IRES 1 (149..245) | 239 | chr10 (+): 98163388 | 0 |
| IRES 1 (149..245) | 240 | chr10 (+): 98163389 | 0 |
| IRES 1 (149..245) | 241 | chr10 (+): 98163390 | 0 |
| IRES 1 (149..245) | 242 | chr10 (+): 98163391 | 0 |
| IRES 1 (149..245) | 243 | chr10 (+): 98163392 | 0 |
| IRES 1 (149..245) | 244 | chr10 (+): 98163393 | 0 |
| IRES 1 (149..245) | 245 | chr10 (+): 98163394 | 0 |
Experimentally-Probed TIS (eTIS)
| Interconnected IRES | TIS Transcript Location |
TIS Chromosomal Location |
Codon |
|---|
Predicted Non-canonical TIS (nTIS)
| Interconnected IRES | TIS Transcript Location |
TIS Chromosomal Location |
Non-canonical Start Codon |
Confidence |
|---|
uORF
| Interconnected IRES |
uORF Transcript Location |
uORF Chromosomal Location |
uORF Sequence | Sequence |
|---|---|---|---|---|
| IRES 1 (149..245) | 66..95 |
chr10 (+) : 98162898 - 98162927. |
30 bps (seq) | AUGUUGUCCAGGCUGGUGUCGAAAUCCUGA |
| IRES 1 (149..245) | 204..212 |
chr10 (+) : 98163353 - 98163361. |
9 bps (seq) | AUGUUGUAG |
| IRES 1 (149..245) | 235..471 |
chr10 (+) : 98163384 - 98163397, chr10 (+) : 98231512 - 98231687, chr10 (+) : 98234446 - 98234492. |
237 bps (seq) | AUGUUACUUACAUGUUAUCAGGGAAUACCAAGAGCAGAGAGAGCAUCCAGGAACCUAGAUCUGAUUACUACAAUCAUGAAGUUCCUGAUAUUGACCUCAGUGAUUGUGAAUUCCCACAUGUCAUUGAAAUUUAUGACUUUCCCCAAGAAUUUCAUACUGAAGACCUUCUACGGGUUUUCUGCAGUUAUCAAAAGAAAGGAUUUGAUAUUAAAUGGGUGGAUGAUACACAUGCCCUAG |
ITAF Binding Events
| Interconnected IRES |
ITAF Type | ITAF Transcript Location |
ITAF Chromosomal Location |
ITAF Sequence |
Sequence | Identified Score |
|---|---|---|---|---|---|---|
| IRES 1 (149..245) | eIF4A3 (HITS CLIP, Hela cells replicate 1) | 144..165 |
chr10 (+) : 98163293 - 98163314. |
22 bps (seq) | GUGAGGCUGCUGUCAGAAAGGA | 3 |
| IRES 1 (149..245) | eIF4A3 (HITS CLIP, Hela cells replicate 2) | 155..237 |
chr10 (+) : 98163304 - 98163386. |
83 bps (seq) | GUCAGAAAGGAACUUUUACACAGUUUGCCUUCAGAGACAGUCUAUCGGCAUGUUGUAGAGUUUUAUUACUAAGAAAAUAAAUG | 8 |
| IRES 1 (149..245) | eIF4A3 (HITS CLIP, Hela cells replicate 1) | 181..248 |
chr10 (+) : 98163330 - 98163397. |
68 bps (seq) | GCCUUCAGAGACAGUCUAUCGGCAUGUUGUAGAGUUUUAUUACUAAGAAAAUAAAUGUUACUUACAUG | 42 |
IRES Activity Measurements
| Interconnected IRES |
IRES Activity |
Promoter Activity |
Splicing Activity |
Matched Transcript Location |
Matched Transcript Sequence | Matched Chromosomal Location |
Designed Human Oligo Sequence |
Designed Oligo Sequence | BLASTN Query Cover |
BLASTN Percent Identity |
|---|
IRES Elements Verified in the Literature
| Interconnected IRES |
Matched Transcript Location |
Matched Transcript Sequence | Matched Chromosomal Location |
Literature Verified Sequence |
Verified Sequence | BLASTN Query Cover |
BLASTN Percent Identity |
Reference |
|---|
Download
The track data of this page can be download via the following two formats:
- Copyright © 2020, NUK All rights reserved. Computational Biology & Intelligence System Lab